Assessment of the antibacterial activity of an anti-infective molecule on persister cells
In vitro/ex vivo testing Antibacterial and antifungal
Background of persisters model
‘Resistance’ describes the inherited ability of microorganisms to grow at high concentrations of an antibiotic, irrespective of the duration of treatment, and is quantified by the minimum inhibitory concentration (MIC).
Molecular mechanisms of resistance include mutations in the drug target, enzymatic activity that directly inactivates the antibiotic and the activation of efflux pumps.
Bacteria are also able to survive extensive antibiotic treatments without acquiring resistance mutations: ‘tolerance’ and ‘persistence’.
‘Tolerance’ describes the ability of microorganisms to survive transient exposure to high concentrations of an antibiotic without a change in the MIC.
In contrast to resistance and tolerance, which are attributes of whole bacterial populations, ‘persistence’ is the ability of a subpopulation of a clonal bacterial population to survive exposure to high concentrations of an antibiotic.
The frequency of persister cell formation is high during stationary-phase, stress conditions, and biofilms states. Since bacteria often encounter these conditions during host infection, persisters are clinically relevant as contributors to chronic and relapsing infections as well as the development of antibiotic resistance.
Objectives of the persisters model
To determine the anti-persisters activity of a drug in development using persister cells from Acinetobacter baumannii and Pseudomonas aeruginosa.
Our approach at Vibiosphen
• The A. baumannii and P. aeruginosa strains used are susceptible to tigecycline and ciprofloxacin, respectively.
• Bacteria were grown to obtain stationary phase cells.
• Stationary phase cultures were washed and treated with 100 × MIC tigecycline or ciprofloxacin for 4 h.
• Bacteria were then washed and treated with a range of concentrations of the test compound or control antibiotics.
• At specific time points a sample was removed, serially diluted, and plated to determine viable cell counts (cfu).
Outcomes of persisters model
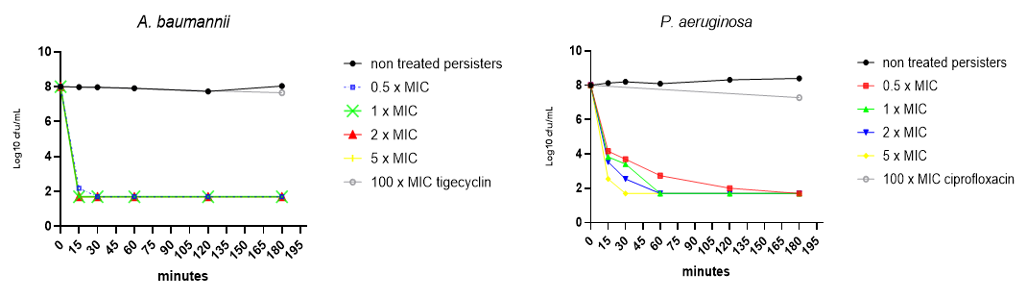 Figure. Killing kinetics of test compound against persister A. baumannii and P. aeruginosa cells. LLOD=50cfu = 1.69 log10
Figure. Killing kinetics of test compound against persister A. baumannii and P. aeruginosa cells. LLOD=50cfu = 1.69 log10
• Incubation with the test compound resulted in the death of A. baumannii and P. aeruginosa persister cells within minutes, at a concentration as low as 0.5 x MIC.
Why Choose Vibiosphen?
• Proven expertise in preclinical research
• Flexible and customized study designs adapted to sponsor requirements (see downloads available as example)
• Advanced facilities to ensure reliable and reproducible results (BSL2 and BSL3)
• Strong collaborations with pharmaceutical companies, biotech firms, and academic partners
Vibiosphen combines scientific excellence with industry know-how to deliver actionable preclinical data that drive innovation in infectious disease therapeutics.
Contact Us
If you are developing new treatments or vaccines, Vibiosphen can help accelerate your research.
Contact us today to discuss your project and explore how our Pharmacokinetics models can support your development strategy.
We will be pleased to facilitate your project by providing a customized study design to your project objectives.
